Introduction¶
This documentation gives a short introduction to Chemaxon's Trainer Engine.
What is Trainer Engine?¶
Chemaxon's Trainer Engine is a tool to predict molecular properties by training machine learning models on input data sets. This tool supports the model life cycle management with:
Data preparation¶
- Normalization of chemical structures (standardization)
- Transformation of molecules to descriptors (feature generation)
Model training¶
- Fitting various model types on the input descriptors and labeled data
- Calculation of statistics to measure model accuracy
Optimization and validation¶
- Visualization and comparison of model details
- Provides a central repository of training and prediction runs for reproducibility
Deployment¶
- Integration end-points for model building and inference
- Makes predictions on novel molecules interactively or in batches
- Enrichment of predicted values with applicability domain information and error prediction
What is the benefit of using the Trainer Engine?¶
Trainer Engine translates input data into executable predictions. It has been used to build successful models for wide range of measured data types including:
- phys-chem properties (e.g. boiling point, vapor pressure, logP, logD)
- analytical chemistry data (retention time)
- ADMET end-points (PAMPA, Caco2 permeability, hERG, MetStab, BBB penetration, CYP inhibition, PAINS)
- on-target assay end-points (different target families includes receptors like GPCRs, enzymes e.g. kinases and transporters).
Trainer Engine predictions are supporting medicinal and analytical chemists, toxicologists, drug discovery project team members to assess risks and opportunities of the compound collections.
It enables computational chemists to experiment with different model types, explain their behavior and seamlessly provide access to high-quality models for larger audiences.
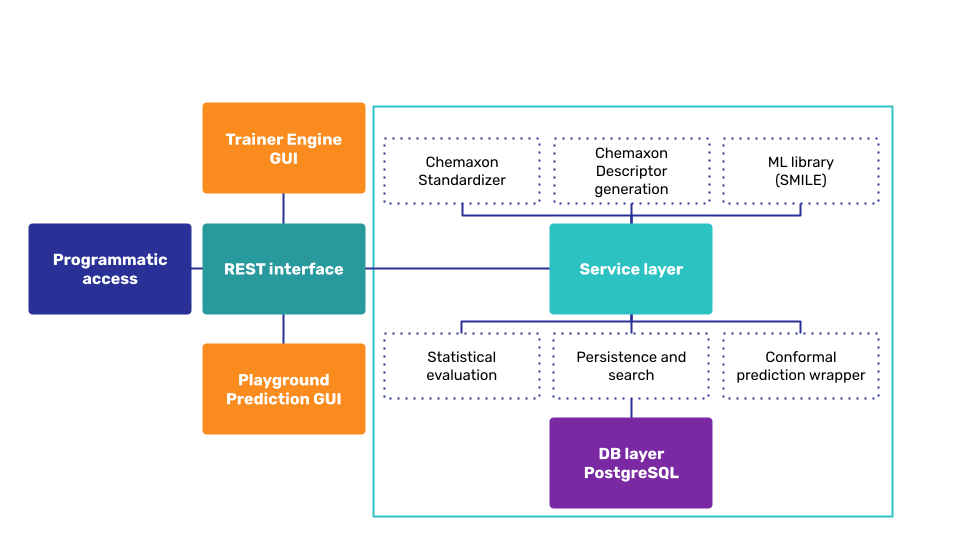
Overview of the components¶
Trainer Engine is a service application interfaced with
- Trainer web user interface (GUI) for model building, assessment, optimization, analysis and management workflows
- REST API end point with SWAGGER API docs for integration
- Default integration with an interactive single or batch prediction web interface, called Playground
The trainer service includes Standardizer, Descriptor generation, ML model building libraries. It calculates the statistical values and implements multi-step workflows, like conformal prediction. All input molecules, configurations, calculated values and statistics are stored in a persistency layer in a PostgreSQL database.
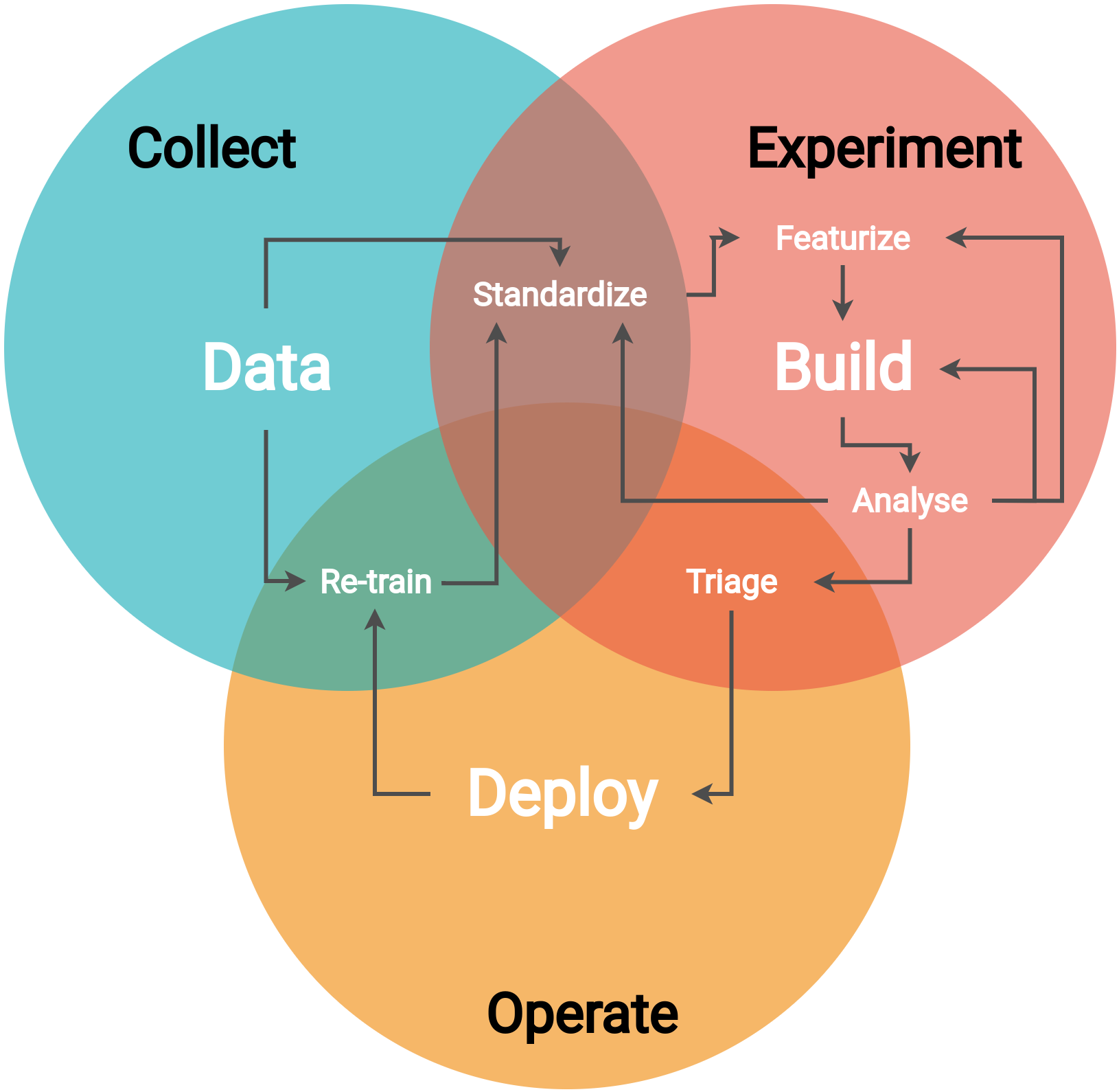
Overview of the general workflow¶
The general machine learning workflow consists of the following steps:
- Upload the input file (sdf) containing molecules and labeled data
- Configuration of standardization actions and descriptor generation
- Configuration of training (model building) and validation via split
- Model building, that can be extended with configurable applicability domain assessment and error prediction
- Automatic model assessment on the training set and test set
- Visualization of the accuracy metrics, model details (feature importance values) and relationships between chemical structures and prediction accuracy
- Optimization of descriptor generation and training parameters (iteration from step 2)
- Selecting the best scored models and setting "In production" flag
- Predict new molecules in single or batch mode with Playground interface
- Integrate production models into design workflows using the REST interface
Programmatic interaction¶
The REST API end points offer all utilities to ingest data, train models and get statistical assessment results. This extension capability allows automatic parameter optimization and selection of the best performing models. API docs is provided via SWAGGER, proof of concept Jupyter notebooks covering larger workflows are available on demand. Contact us for further details: calculators@chemaxon.com.
Design ecosystem¶
Seamless early stage discovery project and hypothesis management ecosystem is offered through integrating Trainer Engine with Chemaxon's Design Hub. In this setup the built models are made available as Design Hub plugins supporting triage of hypothetical molecules and fostering multi parameter optimization when ideas are assessed based on the most important attributes delivered by Trainer Engine and the additional plugins available in Design Hub. Throughout the Trainer Engine API interface, models can be asynchronously built or re-trained and deployed into Design Hub.