Chemaxon's hERG Predictor¶
This documentation gives a short introduction to Chemaxon's hERG Predictor.
Table of Contents¶
- What is hERG?
- Chemaxon's hERG models
- How to use the hERG Predictor?
- Licensing
- References
- Release notes
What is hERG?¶
hERG potassium channels play an essential role in normal electrical activity of the heart, mediating the cardiac action potential of the heartbeat. Affecting their activity by xenobiotics can have life-threatening consequences. Accordingly, hERG is one of the most important off-targets of drug discovery.
Optimisation to reduce the risk of inhibiting the hERG channels during discovery projects requires computational prediction in the early design throughout the pre-synthesis phase. The hERG channel inhibition capacity of a drug is measured by its hERG activity (Act).
Experimentally, hERG activity is determined by different electrophysiological methods and measured as IC50 or Ki values. However, as with other physico-chemical properties (e.g. pKa), the negative logarithm of the measured activity (pActivity) is used in the literature:
pActivity = -log10(Act)
Besides quantitative hERG values it is also common to provide a two-class classification of compounds for hERG activity.
Chemaxon's hERG models¶
We built two hERG models, an activity model and a two-class (TOXIC or SAFE) classification model.
The hERG activity model¶
The activity model was built using multiple publicly available data sources, including the hERG Central DataBase, ChEMBL and various selected publications and patents. The current version of the activity hERG model encapsulates structure activity relationships (SARs) from around 2500 pActivity data points.
We ran a test to validate the predictive capabilities of the activity model on a test set of around 270 molecules. The test resulted in a 0.80 Pearson correlation coefficient and a 0.55 RMSE. 72% of the test set molecules had delta distribution less than 0.5.
The accuracy report is available here.
The hERG classification model¶
The classification model was built using the Honma et al. dataset. The current version contains around 204k training data points.
The classification classes of the model were determined based on the IC50 threshold of 10 μM. TOXIC corresponds to predicted IC50 ≤ 10 μM or ≥50% inhibition at 10 μM, while SAFE corresponds to predicted IC50 > 10 μM or <50% inhibition at 10 μM.
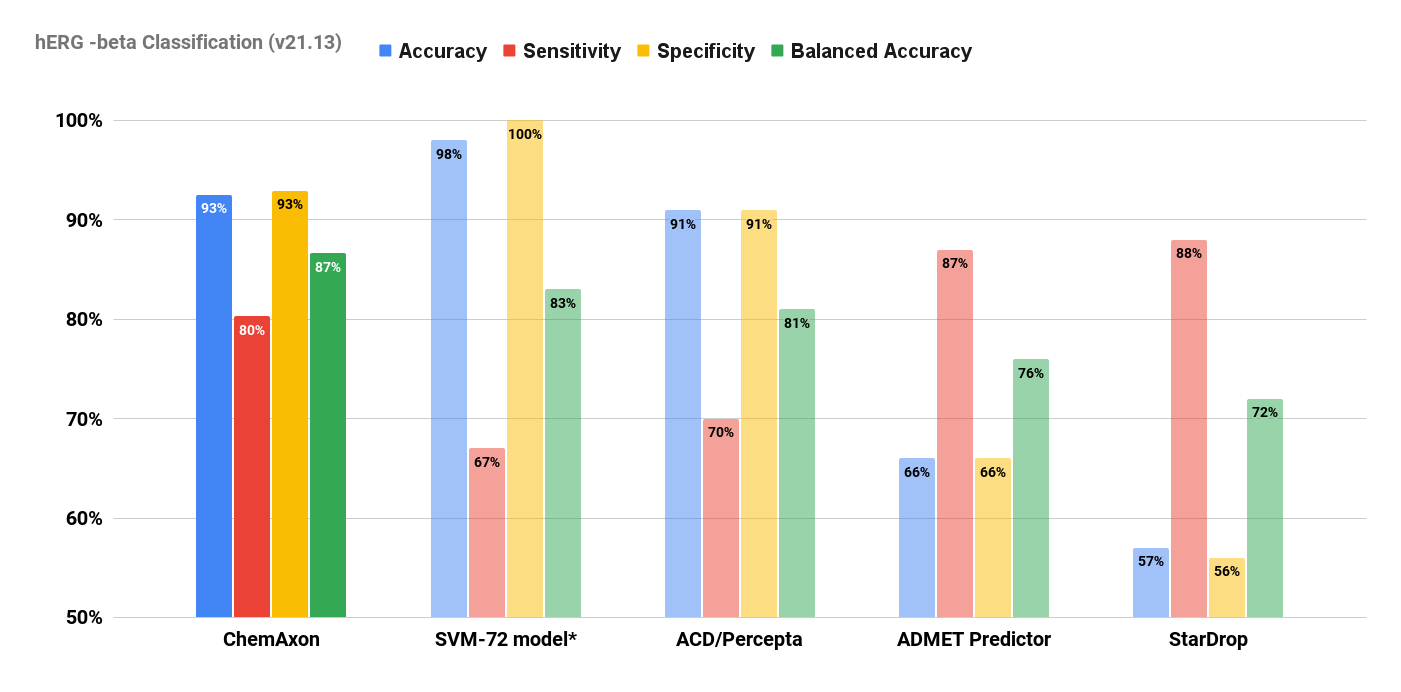
We ran a test to validate the predictive capabilities of the classification model on the remaining test subset of around 87k data points.
Our results were compared to the performance of other commercial hERG models in the article. They are summarised below.
**_NOTE:_** The hERG classification model is currently **ONLY** available in [MarvinSketch](#marvinsketch), [cxcalc](#cxcalc) and [Chemical Terms](#chemical-terms).
Model creation and applicability domain¶
For both models we used Random Forest and Conformal Prediction algorithms to create them and provide the applicability domain of the prediction. The applicability domain provides information about a model's performance and accuracy.
For creating the models we used selected fingerprints (e.g. ECFP) and physico-chemical descriptors.
The applicability domain is based on the 5 most similar compounds from the training set according to ECFP-4 fingerprints and Tanimoto metrics. The error bound of the applicability domain comes from the Conformal Prediction algorithm.
**_NOTE:_** The hERG applicability domain is currently **ONLY** available in [Design Hub](#designhub).
How to use the hERG Predictor?¶
Chemaxon's hERG Predictor uses the activity and classification models to predict the pActivity value and the classification class.
It is currently available in MarvinSketch, in the Design Hub web application, in the cxcalc command line tool and the Chemical Terms language.
MarvinSketch¶
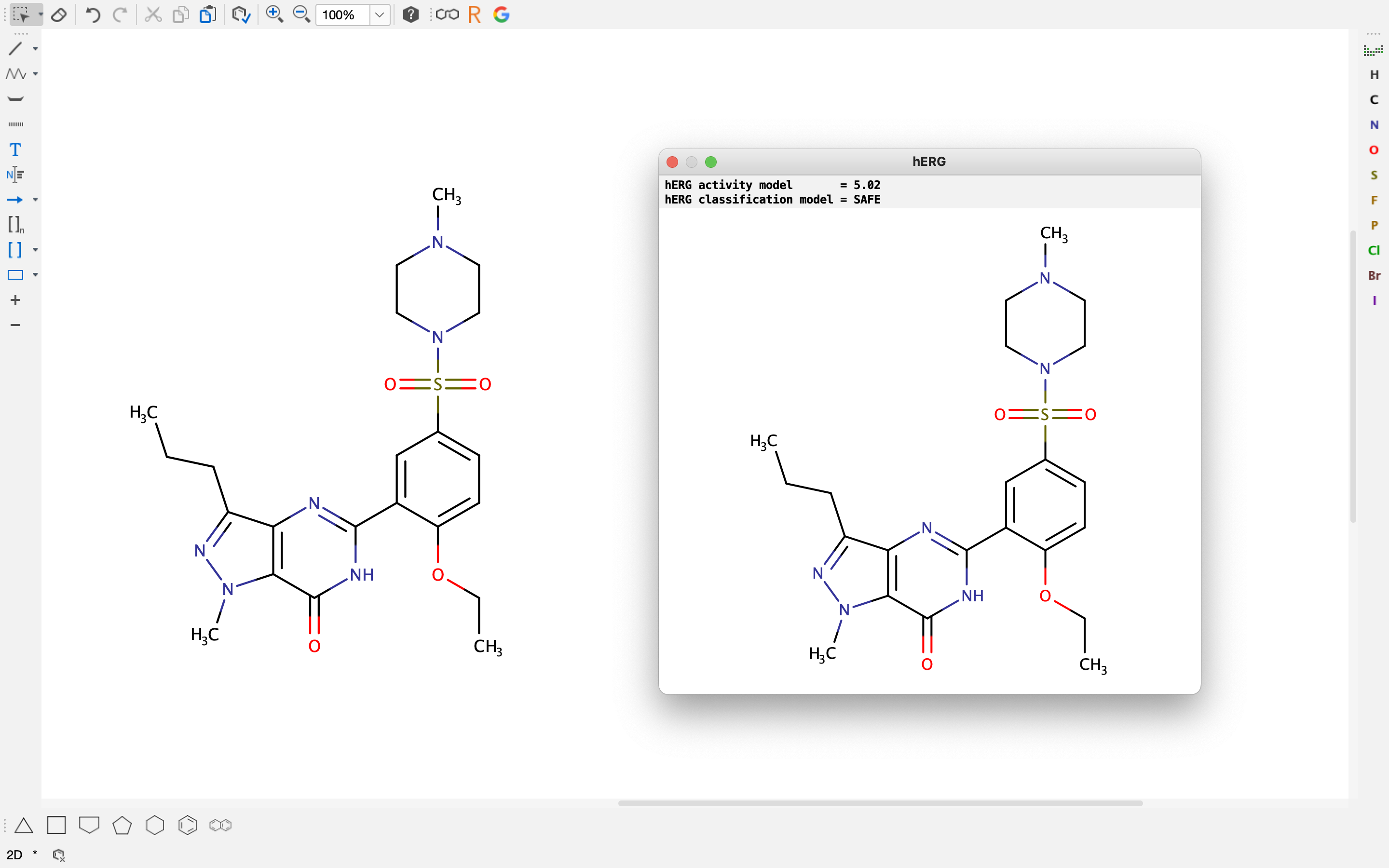
In MarvinSketch hERG prediction can be done with the hERG Plugin, which can be found under the Calculations » ADMET menu item. The current version of the plugin returns the hERG activity and the classification class as well.
Design Hub¶
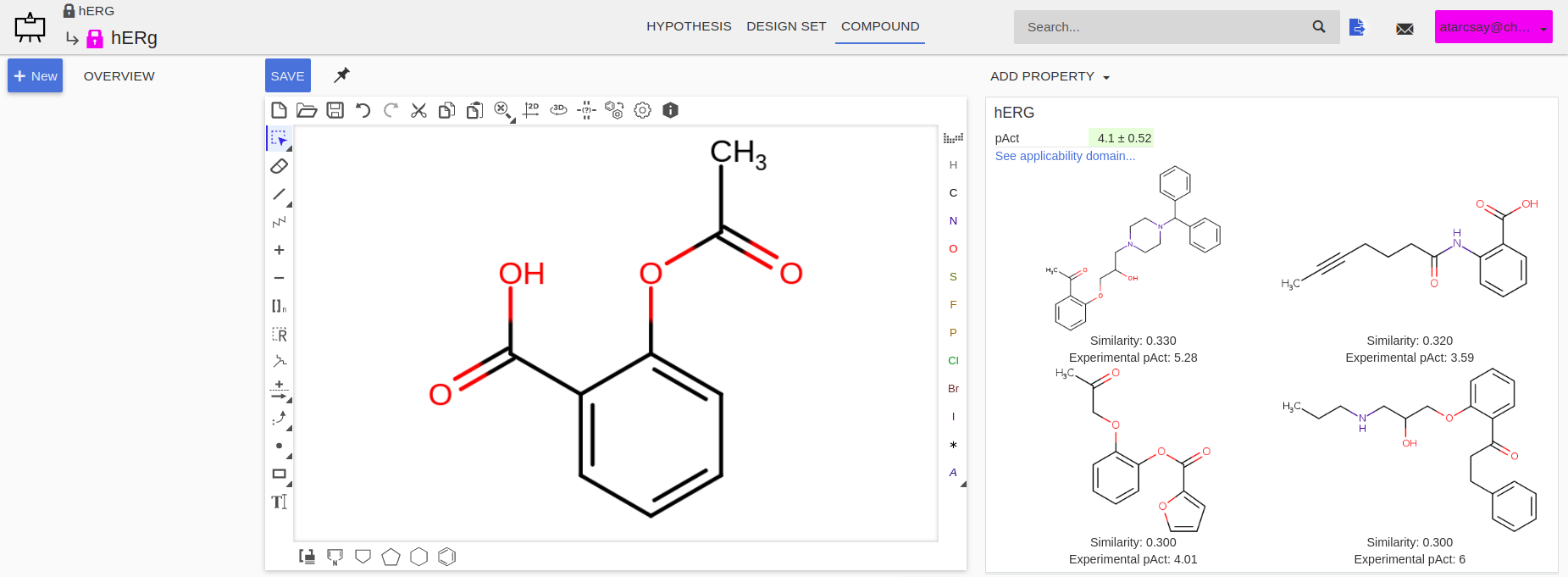
In Design Hub predicting hERG activity can be done with the Add Property » hERG menu item. The predicted activity value with its applicability domain appears next to the drawing canvas.
cxcalc¶
To predict hERG with cxcalc, use the herg function. You can choose the model for prediction by setting the -a (for the activity model) and the -c (for the classification model) options. The default setting is true for both options.
The following examples show how to use cxcalc for hERG prediction:
Chemical Terms¶
To predict hERG with Chemical Terms use the herg() or hergActivity() functions for activity prediction and the
hergClass() function for classification class prediction.
The following examples show how to use Chemical Terms for hERG prediction:
**_NOTE:_** Memory issues (slowdown, prediction failure) can be experienced when predicting hERG in Instant JChem (IJC). To overcome such issues, increase the heap memory available to IJC. For more information see the relevant [memory management documentation](https://docs.chemaxon.com/display/docs/instant-jchem_memory-usage.md) page.
Java API¶
Since v. 23.17 the hERG activity and classification class predictors are also available via the public Java API. You can find their API
documentation pages here and here.
JChem Microservices¶
The hERG activity and classification class predictors are also available via the JChem Microservices. You can find their documentation here and here.
Summary table¶
The following table summarises the availability of the hERG prediction and applicability domain features in the above mentioned products.
| Product name | hERG prediction | Applicability domain |
|---|---|---|
| MarvinSketch | ✅ | ❌ |
| Design Hub | ✅ | ✅ |
| Cxcalc | ✅ | ❌ |
| Chemical Terms | ✅ | ❌ |
| Java API | ✅ | ❌ |
| JChem Microservices | ✅ | ❌ |
Licensing¶
To access and use the hERG Predictor you need a valid Chemaxon ADMET license. Please consult us for more information on licensing.
References¶
- Du, Fang et al. (2011). hERGCentral: A large database to store, retrieve and analyse compound-human ether-à-go-go related gene channel interactions to facilitate cardiotoxicity assessment in drug development, Assay and Drug Development Technologies, 9(6), 580-588. DOI link
- ChemBL 27, ChemBL database
- Honma, Teruki et al. (2019). Support Vector Machine model for hERG inhibitory activities based on the integrated hERG database using descriptor selection by NSGA-II, Scientific Reports (NatureResearch), 9, 12220.
DOI link